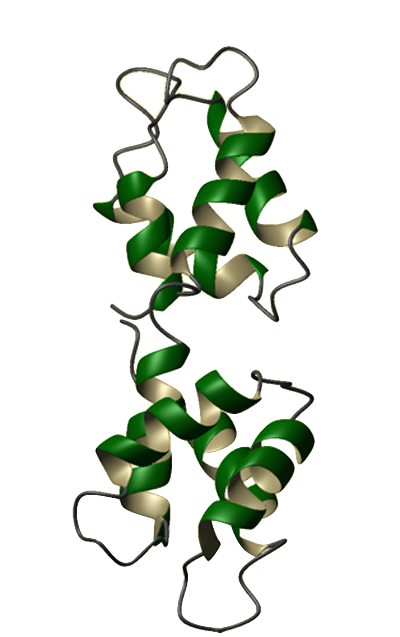BMRB Featured System: Calmodulin |
|||||
| |||||
Eukaryotes utilize calcium ions to regulate a wide range of cellular processes via intracellular signaling pathways. The generic signaling scheme typically begins with an external signal. This signal causes an increase in a cell's calcium concentration. Regulatory proteins in the cytoplasm recognize the change in concentration and consequently bind calcium ions. The calcium-bound regulatory proteins are then able to bind specific targets, thus relaying the signal to the next pathway constituent. One of the best known examples of such a regulatory protein is calmodulin (CaM). Calmodulin is a calcium modulator protein and a transducer of calcium signals. It has a molecular mass of 16,790 Da and contains 148 amino acid residues. Abundant in all eukaryotic cells, CaM's sequence has been highly conserved throughout evolution. The amino acid sequences of CaM in animals, plants, fungi, and protozoans differ from each other by only a few dozen residues. Humans even share identical CaM sequences with many other animal species, including echinoderms, fish, frogs, birds, and other mammals. Calcium-free CaM (apo-CaM) is generally a dumbbell-shaped protein, composed of 2 globular (calcium-binding) domains joined by a flexible connector. Each globular domain contains a pair of high-affinity calcium binding sites known as EF-hands. The connector is a long alpha helix that separates the globular domains. Calcium-bound CaM (Ca2+-CaM) can assume a variety of shapes depending on the target. Our understanding of calcium-binding proteins began in the 1950s and 60s when Setsuro Ebashi and his colleagues conducted a series of studies on a calcium-binding protein that was later named troponin. The discovery of troponin paved the way for the rapid discovery of other calcium-binding proteins, including CaM. CaM was discovered almost concurrently in 1970 by two separate labs investigating cyclic 3',5'-nucleotide phosphodiesterase. It was first described by Wai Yiu Cheung and then by Shiro Kakiuchi and his colleagues later that same year. Cheung found that purified cyclic 3',5'-nucleotide phosphodiesterase exists in a largely inactive form unless supplemented with an unknown activator. Kakiuchi's lab detected the presence of this activator (phosphodiesterase-activating factor) in brain extracts and found that the activity of brain phosphodiesterase is stimulated by µM concentrations of Ca2+. The mechanism by which this "activator" affected phosphodiesterase activity was unknown until the discoveries of Tian Seng Teo and Jerry Hsueh-Ching Wang. In 1972 Teo, Wang and colleagues found that cAMP slowly activates phosphodiesterase, possibly by enhancing the interaction between the enzyme and the activator. A year later, they demonstrated that the activation of phosphodiesterase requires the simultaneous presence of Ca2+ and the activator, and that this activator is a Ca2+-binding protein. By 1978, this activator was found to affect the activities of many other proteins and was commonly known as the calcium-dependent regulator (CDR) or calcium-dependent modulator (CDM). In 1979, this modulator protein was dubbed calmodulin, the widely accepted name today. CaM was chosen as the Protein of the Month for August 2003 by both The Protein Data Bank and the European Bioinformatics Institute. These sites offer additional information pertaining to CaM structure and genomics. A CaM site is maintained by The EF-Hand Calcium-Binding Proteins Data Library at Vanderbilt University. It includes a collection of sequence, structural, functional, and other information about CaM and its role in signaling. In addition, Mitsu Ikura maintains a Cellular Calcium Information Server at the Ontario Cancer Institute. | |||||
|
Next: CaM's function |